
| Theme: Artificial & Business Intelligence | |||
 |
||||||
| Automatic Segmentation of the Left Ventricle in 3-D Echocardiography Images |
 |
|||||
|
||||||
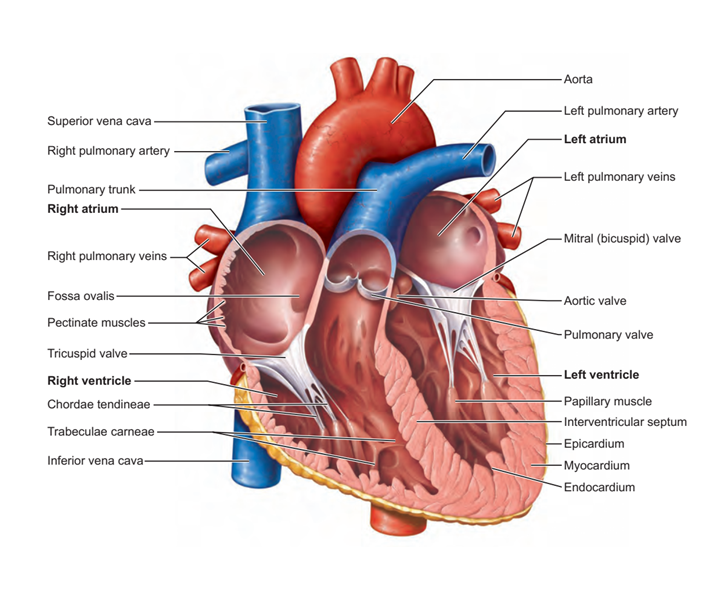
Figure 1: A detailed heart anatomy
The measurement of the left ventricle (LV) volume requires accurate delineation to separate the LV from the adjacent structures. LV manual segmentation can be performed, but it is tedious, time-consuming and vulnerable to intra/inter-observer variability . As a result, deriving quantitative metrics such as ejection fraction (EF), end-diastolic volume (EDV) and end-systolic volume (ESV) is difficult. It is noteworthy to mention that there are some existing semiautomatic segmentation approaches used in medical imaging such as deformable and statistical models, but they suffer from the inability to generalize to different cardiac anatomies outside of the training datasets, in addition to suffering from low accuracy and weak robustness. The incorporation of deep learning (DL) to fully automatically segment the LV volume could have a great impact on improving echocardiography segmentation . It offers a fast segmentation process, allows clinicians to compute extra quantitative clinical analysis, and reduces inter/intra observability to improve and accelerate the diagnosis and the treatment of various cardiovascular diseases.
- Segmentation of 72 LV myocardium and blood pool in ES frame was performed in addition to 5 LV blood pool segmentation in ED frame.
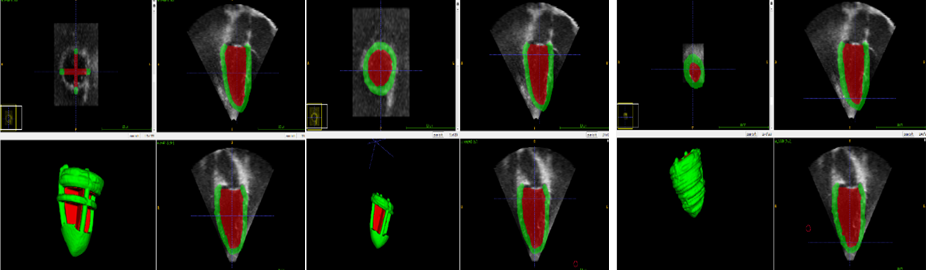
Figure 2: The segmentation steps
- Three pre-processing tasks were performed: (1) intensity normalization, (2) image cropping, and (3) image downsampling.
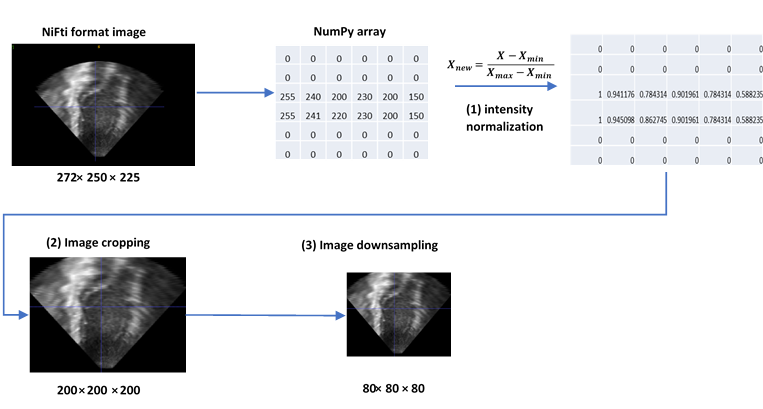
Figure 3: Processing steps
- Data augmentation was performed in order to get a large-scale dataset.
- Then, V-net and U-net were used to train the datasets using five cross-validation techniques to validate the performance of the model during the training.
- The cross-entropy loss and dice similarity coefficient were used to track the performance of the model during the training.
- The best of each model was used to evaluate unseen test images, which included five pathological cases (cardiac hypotrophy) in both ED and ES frames.
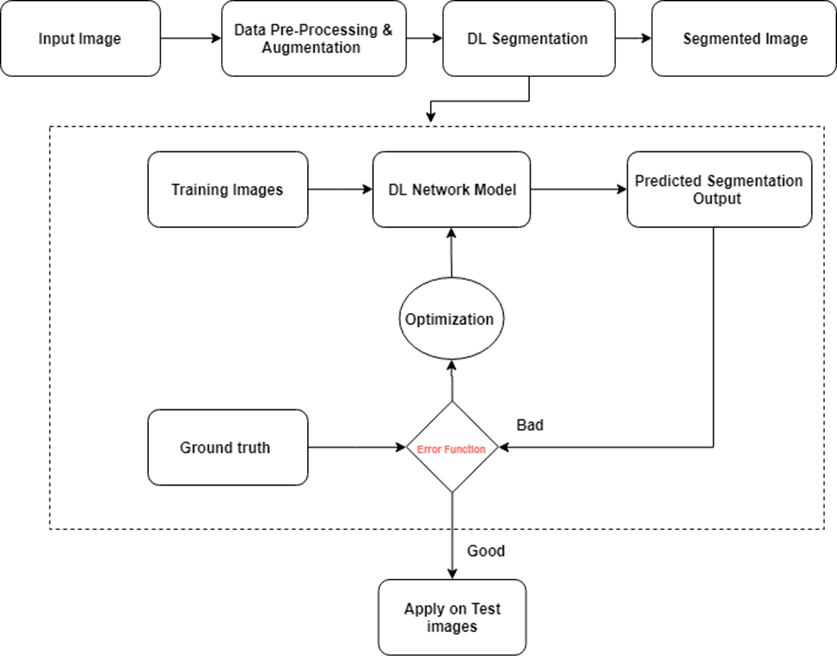
Figure 4: Project Scheme
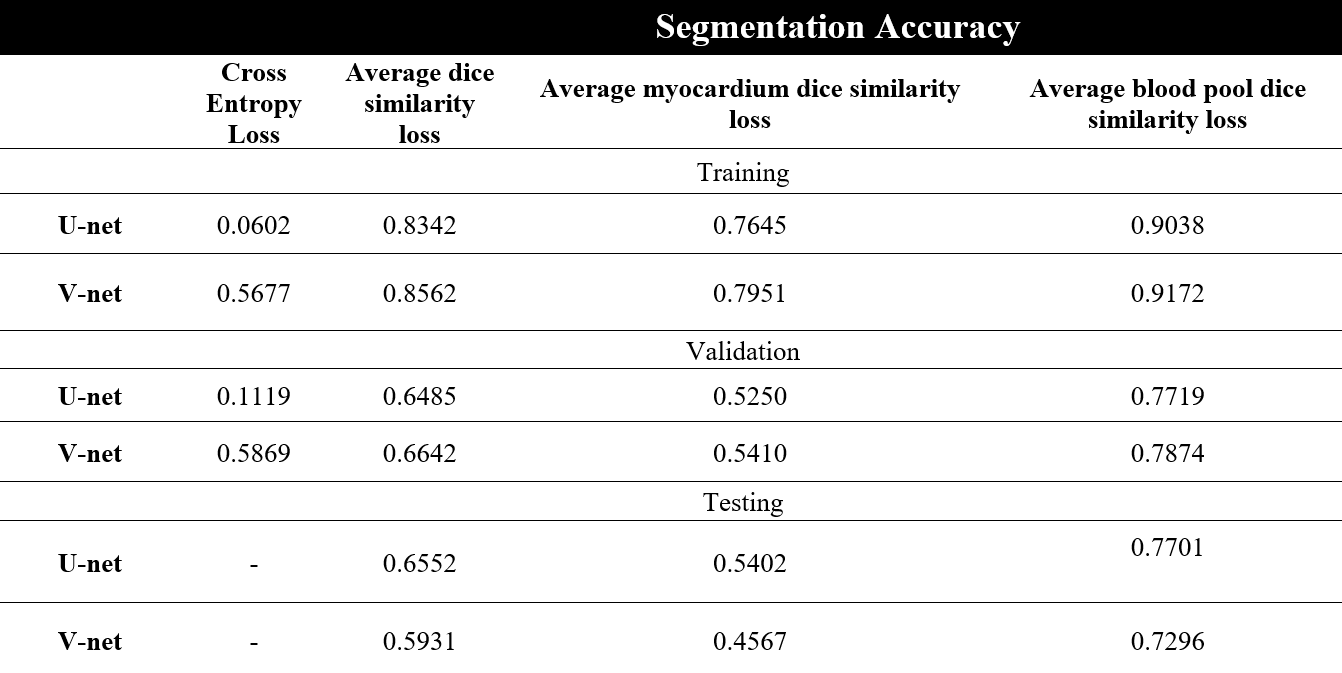
Table 1 : The comparison between U-net and V-net performance
The results of both models were very close to each other on validation sets, but they were not on test sets.
The results of applying DL model on unseen datasets provided an average accuracy of 0.6552 for U-net and 0.5931 for V-net.
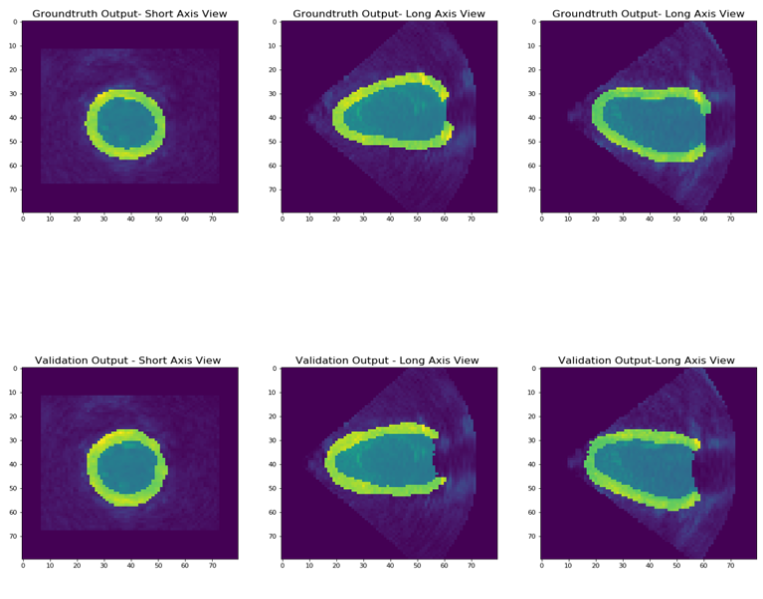
Figure 5: Segmentation output of DL for one subject in all 3D views.
Generally, our findings performed well on unseen data sets with an average accuracy of 0.6552 for U-net and 0.5931 for V-net. Future work should carefully consider the potential effects of :
- Model weight initialization methods
- Manual segmentation accuracy of ES images (particularly myocardium)
- Data augmentation technique and interpolation methods
- Run-to-run variations.
Considering all these factors would definitely contribute to the success of this work, thereby enabling the potential adoption of the method in routine clinical use. This would help clinicians to perform clinical quantitative analysis, which is currently difficult because of the long time required.
I would like to express my gratitude to :
- My clinical supervisor Dr Esther Puyol.
- My academic supervisor Dr Andrew King at King's college London.
- Ezequiel Ortiz a BSc student for his help to understand his project about segmentation in 2D echocardiography.
- Nicholas Byrne for his valuable help to share his implemented V-net network to be used in this project
- Dr Alberto Gomez Herrero for dedicating some time to help me doing manual segmentation.
[1] P. Peng et al., "A review of heart chamber segmentation for structural and functional analysis using cardiac magnetic resonance imaging," vol. 29, no. 2, pp. 155-195, 2016.
[2] M. Avendi, A. Kheradvar, and H. J. M. i. a. Jafarkhani, "A combined deep-learning and deformable-model approach to fully automatic segmentation of the left ventricle in cardiac MRI," vol. 30, pp. 108-119, 2016.
[3] M. Alsharqi et al., "Artificial intelligence and echocardiography," vol. 5, no. 4, pp. R115-R125, 2018.
[4] P. V. J. a. p. a. Tran, "A fully convolutional neural network for cardiac segmentation in short-axis MRI," 2016.

 Send Email
Send Email